A cell selection tool for spatial proteomics powered by combinatorial optimization.
Modern spatial proteomics techniques, such as Deep Visual Proteomics, depend heavily on high‑precision laser‑capture microdissection at the single‑cell level to isolate cells of interest for proteomics analysis. CellPick seamlessly integrates into these workflows by strategically selecting cells from any tissue region, and by providing precise spatial metadata relative to defined landmarks, accelerating research throughput and enhancing reproducibility.
strategic Cell selection
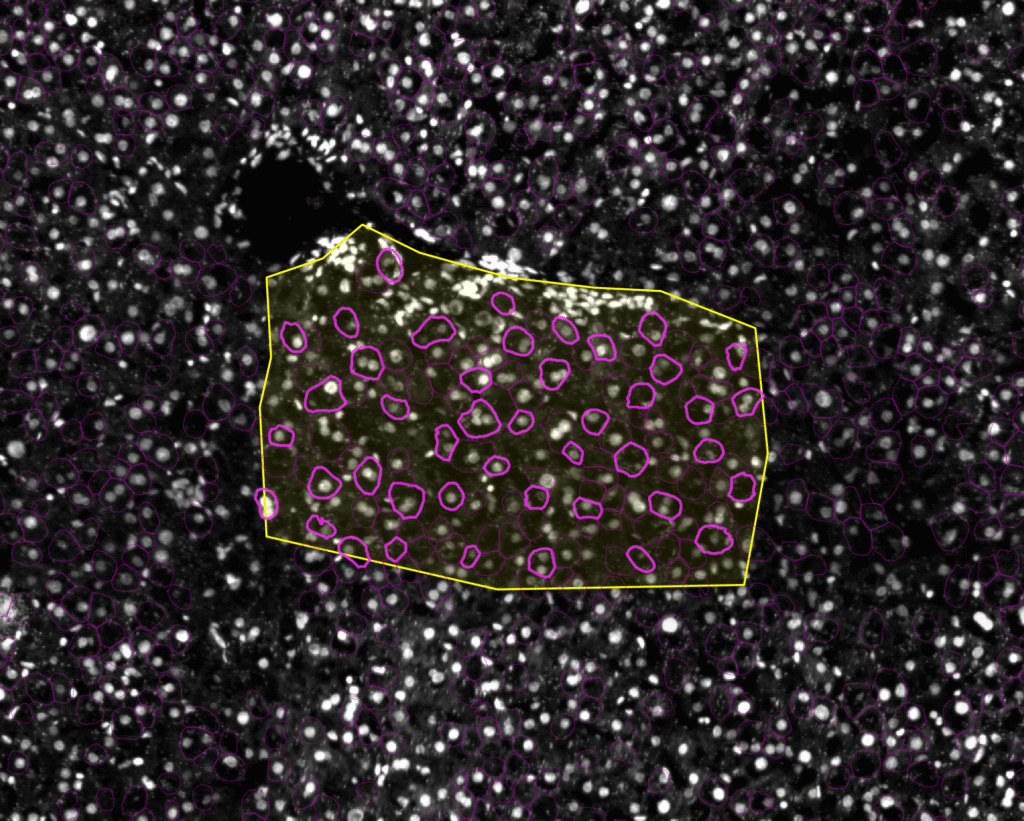
Laser microdissection enables the precise isolation of individual cells from tissue sections using a high-powered laser. While random or manual selection strategies may suffice in regions with a high number of cells, they often lead to the selection of contiguous cell groups in sparsely distributed areas, raising the risk of cellular damage during dissection.
To address this limitation, CellPick incorporates a custom shape selection method based on combinatorial optimization, ensuring the selection of non-contiguous cells while approximately maximizing coverage within defined tissue boundaries.

automatic spatial annotation
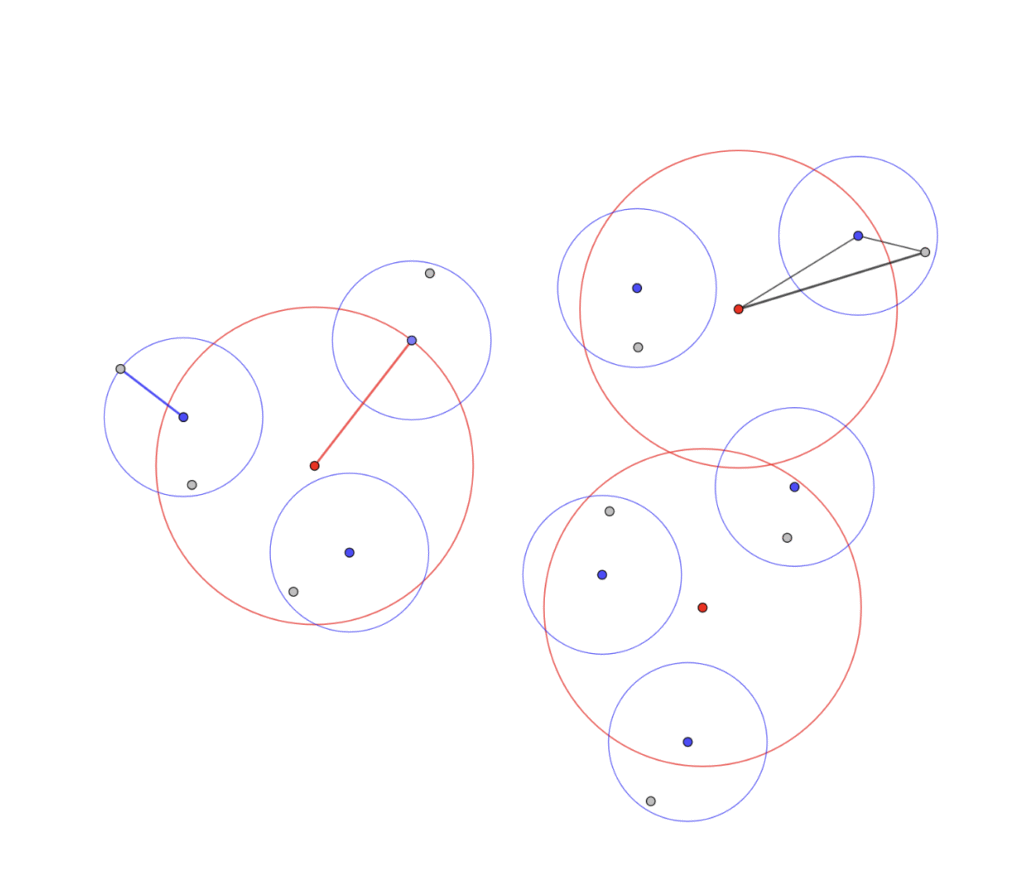
CellPick enables the definition of two user-specified regions of interest, referred to as landmarks, to establish a spatial gradient along a biologically relevant axis, for example, between distinct vein types in the liver. Each selected cell is automatically annotated with a value representing its relative proximity to these landmarks, allowing researchers to position cells along the gradient. This facilitates downstream analysis by enabling correlations between protein expression levels and spatial localization within the tissue.
open source code
Contribute to our code, open issues for feature requests, and download the latest development builds of CellPick.
COming Soon…
read the paper
In the preprint accompanying the software, we describe the details of the combinatorial optimization problems that are involved in the cell selection procedure, as well as the algorithms computing the spatial scores.
Install now
We provide a seamless installation via the pipx package manager. Moreover, we provide a comprehensive documentation, including tutorials on the usage of CellPick, and descriptions of the functions used in the code, allowing for extensibility.
Contact Us
Authors: Paolo Pellizzoni, Lucas Miranda.
Max Planck Institute of Biochemistry
Address
Am Klopferspitz 18
82152 Martinsried, Germany.
lastname@biochem.mpg.de